Antibiotic resistance and Pseudomonas virulence: two sides of the same coin
In people with cystic fibrosis, some resistance to antibiotics caused by Pseudomonas aeruginosa may paradoxically increase bacterial virulence, complicating lung infections and opening the way to new therapeutic approaches.
Mylène Vaillancourt, M.Sc., associate researcher
Peter A. Jorth, Ph.D., Principal Investigator
2021
Cystic Fibrosis (CF) is an inherited and potentially fatal condition characterized by sinopulmonary infections, severe lung damage, as well as disorders in the gastrointestinal system and other organs. In patients with CF, mutations in the CFTR gene cause the absence or dysfunction of the CFTR channel on the surface of the airway epithelium and this interferes with the transport of chloride ions between epithelial cells and the respiratory tract. This ion imbalance affects the water content of the mucus layer on the surface of the respiratory tract, resulting in the accumulation of thick mucus that is conducive to bacterial colonization. One of the most common bacterial species that causes lung infections in CF patients is Pseudomonas aeruginosa [1]. Present in only 10% of infants with CF, its prevalence increases with age to reach 50% in patients over 25 years of age [1], demonstrating its ability to supplant other bacterial species and to chronically infect CF lungs.
P. aeruginosa has a remarkable ability to adapt to various environments. Thanks to this versatility, this bacterium is easily found in natural environments such as soil and water. Unfortunately, this characteristic also makes it a notable opportunistic pathogen that causes serious acute and chronic hospital-acquired infections of the lungs (CF, acute lung injury), skin (burns), and other organs such as the bladder and intestine. To survive and replicate in these different environments, P. aeruginosa changes its behavior. For example, it mainly uses oxygen to produce its energy through aerobic respiration, but can change its metabolism to fermentation if the environment is limited in oxygen. Another important aspect of the versatility of P. aeruginosa is its great mutability under the selective pressure of evolution which gives it new advantageous capacities. An obvious example is the intensive use of antibiotics in patients with CF, who select bacterial isolates that carry mutations in key genes involved in antibiotic resistance.
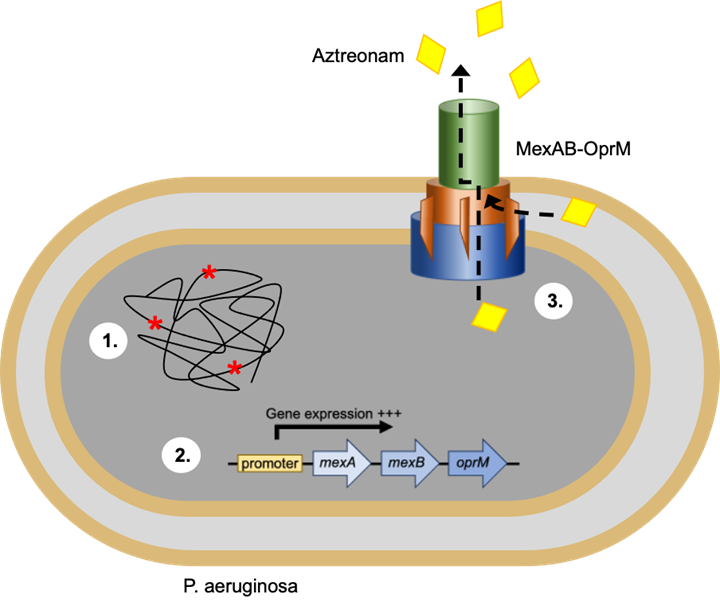
Our team of researchers studied in depth the genetic variability of P. aeruginosa in the lungs of patients with CF as well as the mechanisms that underlie the development of antibiotic resistance in this bacterium. He specifically studied the mechanisms involved in bacterial resistance to aztreonam, the first monobactam antibiotic approved by the U.S. Food and Drug Administration (FDA) for use in patients with CF [2]. Subsequently, the team evolved P. aeruginosa in vitro under continuous and cyclical exposure to aztreonam in order to reflect the different protocols for the administration of antibiotics in patients with CF. The entire genomes of isolates resistant to aztreonam were then sequenced in search of mutations responsible for the development of this resistance. This study revealed two genes that were mutated repeatedly in resistant isolates: NalD and mexR. These genes are both known to participate in the repression of the mexAB-Oprm efflux pump. Efflux pumps are active channels located on the surface of the bacterial cell membrane. Bacteria can use these efflux pumps to expel an antibiotic, thus protecting themselves from the action of that antibiotic. Inactivating mutations in the nALD and mexR genes trigger the overexpression of mexAB-Oprm by P. aeruginosa, making them resistant to aztreonam (Figure 1). This result is particularly important, because, in the literature, mutations in these two genes have been found in 7 to 47% of isolates of P. aeruginosa from CF patients.
During this same study, our group did more than find the mutations responsible for aztreonam resistance. He made comments concerning P. aeruginosa resistant to aztreonam that led to his current research. He first found that the selection of aztreonam could lead to resistance to multiple drugs. For example, isolates resistant to aztreonam also had 2 to 5 times greater resistance to ciprofloxacin. He also observed increased virulence of the Nald and MexR mutants of P. aeruginosa during an acute lung infection in mice. This hypervirulence in P. aeruginosa resistant to aztreonam was surprising, as antibiotic resistance is generally associated with a decrease in virulence and adaptive capacity. This modification of the bacterial phenotype is well known and has clinical implications for patients. For example, the virulent isolates of P. aeruginosa can cause acute infections that trigger an exacerbated inflammatory response and cause lung tissue damage and lung exacerbation in patients. These aggressive isolates are generally sensitive to antibiotics and can be eradicated after appropriate antimicrobial treatment. During acute infections, a small subset of isolates of P. aeruginosa may mutate to acquire some degree of resistance to the antimicrobial, which may result in chronic lung infections. These isolates adapt to the lung environment, lose their virulence and develop a biofilm phenotype. This phenotypic change decreases the inflammatory response and lung tissue damage, but makes these isolates extremely difficult to eradicate due to their antibiotic resistance.
Although this dichotomy between the virulence of P. aeruginosa and its resistance to antibiotics is well known, previous work by our group and other researchers suggests that the two phenotypes are not necessarily mutually exclusive. For example, a recent 5-year prospective study including 510 patients with CF found a significant association between the number of pulmonary exacerbations with hospitalization and the development of aztreonam resistance in P. aeruginosa [3]. This observation could suggest the presence of an isolate of P. aeruginosa which would be both virulent and resistant to aztreonam.
Our current work aims to decipher the mechanisms by which mutations evolved during exposure to aztreonam lead to hypervirulence in P. aeruginosa. More specifically, we are studying the interaction and effects of these hypervirulent strains on immune cells such as monocytes/macrophages. Immunity is an essential factor in bacterial clearance, not only in CF, but in any infectious disease; thus, an altered host immune response can affect the outcome of an infection. A recently published case study nicely highlights the effect of innate immunity on a population of P. aeruginosa during the acute lung infection of a patient in intensive care [4]. In their report, the authors demonstrated how the immune response rapidly decreased the loads of P. aeruginosa during the initial infection, even before the first administration of antibiotics. Les Pseudomonas The remaining ones showed increased sensitivity to antibiotics, which potentiated the action of additional antimicrobial therapy and caused the suppression of the bacterial population.
Monocytes/macrophages are one of the first immune responses to bacterial infection. During early infection, they “eat” infectious bacteria in a process called phagocytosis. Phagocytosis causes macrophages to release antimicrobial mediators that kill pathogens. Macrophages also act as garbage collectors by removing dead cells and debris from the lung, which also helps to reduce inflammation and prevent a disproportionate immune response. Bacteria can develop several different mechanisms to evade phagocytosis and improve their survival in CF lungs. For example, P. aeruginosa uses glycolipid biosurfactants called rhamnolipids to infect epithelial cells and to evade immune cell phagocytosis [5]. Les Pseudomonas Virulent agents can even secrete proteases that degrade antimicrobial agents produced by macrophages and other immune cells, and thus interfere with bacterial clearance. It has been shown that Pseudomonas LasB, an elastase that is part of the type II secretion system, degrades the components of complement C3 and factor B, two critical factors in the processes of recognition of pathogens and phagocytosis by macrophages [6]. Bacteria can also use molecules produced by host cells for their own benefit. For example, itaconate is a metabolite that is produced by macrophages in response to bacterial infection. Its anti-inflammatory properties are essential to restore tissue homeostasis and to prevent a disproportionate inflammatory response. Itaconate also has antimicrobial properties by damaging the integrity of the bacterial wall and inhibiting bacterial growth. However, it has been shown that P. aeruginosa evolves under exposure to itaconate and becomes resistant by altering its metabolism to use itaconate as an energy source and to form biofilms [7].
Because of the importance of macrophages during infection, we are studying how the mutation NalD At P. aeruginosa resistant to aztreonam may affect interactions with macrophages. We use a number of approaches to study how P. aeruginosa Mutants NalD develop during the infection of macrophages, change their metabolism and cause damage. The main approaches we use to answer these questions are fluorescent microscopy and next-generation RNA sequencing.
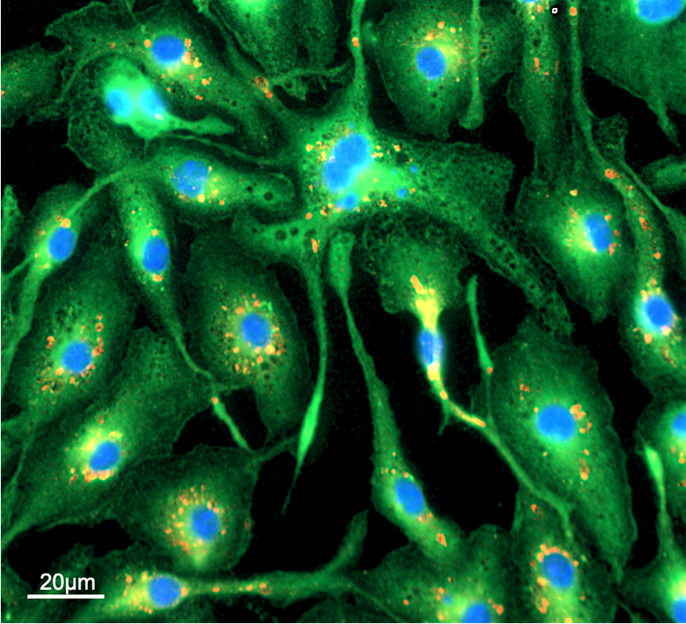
Fluorescence microscopy is a laser-based system that allows the visual analysis of bacteria-macrophage interactions at the molecular level. To perform this technique, the bacteria are incubated with macrophages for determined periods of time, then the infections are preserved chemically before specific molecules on the bacteria and the host cells are labeled with fluorescent antibodies (FIG. 2). These fluorescent antibodies can then be excited by lasers attached to the microscope and highly sensitive detectors capture the fluorescent light emitted by the labeled cells. By marking with fluorescent antibodies in different colors, we can quantify the number of macrophages and bacterial cells, and determine how these cells interact physically with each other. Ultimately, this will help us determine how these bacteria subvert critical immune cells and promote disease progression.
Next-generation RNA sequencing is a technique similar to genome sequencing that allows researchers to determine the complete gene expression profile of a population of cells. Instead of determining the pattern of nucleotide blocks that make up the genome, RNA sequencing is used to count the number of mRNA molecules that are produced from a given gene when its expression is activated. This technique is extremely powerful, as it allows us to determine which of the 6000 genes of P. aeruginosa and 30,000 human genes are activated during infection. This will allow us to identify potential therapeutic targets for these infections that target either the bacteria or the host, based on expression profiles. For example, if P. aeruginosa expresses virulence genes that produce toxins or specific metabolic programs, we will be able to select drugs that specifically inhibit these pathways. Likewise, if macrophages react inappropriately to infection, we can choose drugs that reprogram macrophages to respond in a way that promotes the elimination of P. aeruginosa.
Because of the ability to Pseudomonas To develop multi-drug resistance and to chronically infect CF patients, it is essential to find new molecules with antimicrobial properties that are structurally distinct from current antibiotics. Researchers at the University of Washington have recently begun to explore gallium as a new anti-Pseudomonas therapy. Gallium is different from traditional antibiotics because it is a metal with properties similar to those of iron. It acts like a “Trojan horse” by binding to molecules and proteins that normally fix iron. However, because it reacts differently than iron, it disrupts the iron metabolism of bacteria. Iron is an essential nutrient for the survival and growth of bacteria by acting as a cofactor for a large number of enzymatic reactions. Thus, the substitution of iron with gallium, a metal, reduces the growth and viability of bacteria. An initial clinical trial on 20 patients showed promising results for treatment with gallium nitrate administered intravenously for 5 days to patients with CF [8]. However, a phase 2 clinical study on a larger cohort, the IGNITE study (NCT02354859), did not show a stable change in forced exhalation volume in CF patients, although it did slightly improve respiratory symptoms and decrease bacterial density in cultures of Pseudomonas in the sputum. One of the hypotheses that may explain this lack of clinical effectiveness is the low concentration of gallium measured in patients' tissues after 5 days of intravenous administration. Although gallium concentrations increased in the plasma of CF patients during and a few days after treatments, the concentrations detected were relatively low and barely reached the minimum inhibitory concentration previously established in vitro. The concentrations detected were even lower in sputum. To address this problem, gallium inhalation has been proposed as a more effective delivery method than intravenous administration in a preclinical model [9]. A randomized, double-blind phase 1/2a study is ongoing to test the safety and pharmacokinetics of inhaled gallium in patients with CF (NCT03669614). Because of this promise, we are interested in exploring gallium as a potential therapy to fight infections with P. aeruginosa resistant to antibiotics.
The work of our group and other groups challenges the theory that the acquisition of antibiotic resistance always has a negative impact on the virulence of bacteria. In the specific case of aztreonam, some mutations found in resistant isolates of P. aeruginosa led to 1) multidrug resistance; and 2) increased virulence during macrophage infection. Antimicrobial compounds such as gallium could be an effective treatment in combination with conventional antibiotics to target these “superbugs.”
1. Stephenson A, Wallenburg J, Cheng S, Le T, Le T, Sykes J, Stanojevic S (2020) 2019 Annual Data Report. Cystic Fibrosis Canada. https://www.cysticfibrosis.ca/our-programs/cf-registry
2. Jorth P, McLean K, Ratjen A, Secor PR, Bautista GE, Secor PR, Bautista GE, Ravishankar S, Rezayat A, Garudathri J, Harrison JJ, Harwood RA, Penewit K, Waalkes A, Waalkes A, Waalkes A, Waalkes A, A, A, Singh PK, P, P, R A, Waalkes A, Singh PK, Salipante SJ (2017) Evolved Aztreonam Resistance Is Multifactorial and Can Produce Hypervirulence in Pseudomonas aeruginosa. mBio 8: https://doi.org/10.1128/mBio.00517-17
3. Keating CL, Zuckerman JB, Singh PK, McKevitt M, McKevitt M, Gurtovaya O, Bresnik M, Marshall BC, Saiman L (2021) Pseudomonas aeruginosa Susceptibility Patterns and Associated Clinical Outcomes in People with Cystic Fibrosis following Approval of Aztreonam Lysine for Inhalation. Antimicrobial Agents Chemother 65:e02327-20. https://doi.org/10.1128/AAC.02327-20
4. Wheatley R, Diaz Caballero J, Kapel N, Kapel N, Kapel N, de Winter FHR, de Winter FHR, Jangir P, Quinn A, Del Barrio-Tofiño E, López-Causapé C, Hedge J, Torrens G, Torrens G, Torrens G, Van der Schalk T, Van der Schalk T, Van der Schalk T, Xavier T, Xavier B, Xavier BB, Xavier BB, Xavier BB, Xavier BB, Xavier BB, Fernández-Cuenca F, Fernández-Cuenca F, Arenzana A, Recanatini C, Timbermont L, Torrens G, G, Torrens G, Van der Schalk T, Van der Schalk T, T, Van der A, Ali O, Lammens C, Goossens H, Kluytmans J, Kluytmans J, Kluytmans J, Kumar-Singh S, Oliver A, Malhotra-Kumar S, MacLean C (2021) Rapid evolution and host immunity drive the rise and fall of carbapenem resistance during an acute Pseudomonas aeruginosa infection. Common Nat 12:2460. https://doi.org/10.1038/s41467-021-22814-9
5. McClure CD, Schiller NL (1996) Inhibition of macrophage phagocytosis by Pseudomonas aeruginosa rhamnolipids in vitro and in vivo. Curr Microbiol 33:109 —117. https://doi.org/10.1007/s002849900084
6. Bastaert F, Kheir S, Saint-Criq V, Villeret B, Villeret B, Villeret B, Dang PM-C, El-Benna J, Sallenave J-M (2018) Pseudomonas aeruginosa LasB Subverts Alveolar Macrophage Activity by Interfering With Bacterial Killing Through Downregulation of Innate Immune Defense, Voulhoux R, Sallenave J-M (2018) Pseudomonas aeruginosa LasB Subverts Alveolar Macrophage Activity by Interfering With Bacterial Killing Through Downregulation of Innate Immune Defense, Reactive Oxygen Species Generation, and Complement Activation. Front Immunol 9:1675. https://doi.org/10.3389/fimmu.2018.01675
7. Riquelme SA, Liimatta K, Wong Fok Lung T, Wong Fok Lung T, Fields B, Ahn D, Chen D, Chen D, Lozano C, Sáenz Y, Uhlemann A-C, Kahl BC, Britto CJ, DiMango E, E, Prince A, Prince A (2020) Pseudomonas aeruginosa (2020) Pseudomonas aeruginosa Uses Host-Derived Itaconate to Redirect Its Metabolism to Promote Biofilm Formation. Cell Metabolism 31:1091-1106 .e6. https://doi.org/10.1016/j.cmet.2020.04.017
8. Goss CH, Kaneko Y, Khuu L, Anderson GD, Anderson GD, Anderson GD, Ravishankar S, Ravishankar S, Aitken ML, Lechtzin N, Zhou G, Czyz DM, McLean K, Olakanmi O, Shuman O, Suman HA, Suman HA, Teresi M, HD, Shuman HA, Teresi M, HD, Shuman HA, Teresi M, HD (2018) Gallium disrupts bacterial iron metabolism and has therapeutic effects in mice and humans with lung infections. Sci Transl Med 10:eaat7520. https://doi.org/10.1126/scitranslmed.aat7520
9. Woo J, Hearne K, Kelson A, Yee L, Yee L, Espadas C, Truong VL (2019) 716. In vitro and in vivo Non-clinical Efficacy of AR-501 (Gallium Citrate). Open Forum Infect Dis 6:S322. https://doi.org/10.1093/ofid/ofz360.784
Research
Thanks to Our Partners